How do you fix a primer dimer
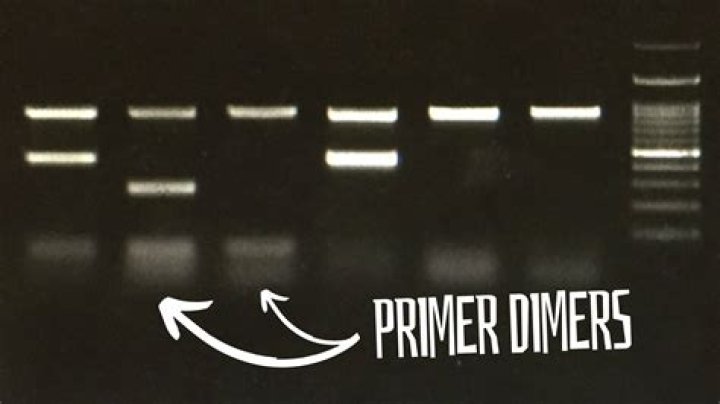
increase the annealing temperature.increase time\ temperature of template denaturation.decrease primers concentration(10 pmol will be OK)use a PCR enhancer such as DMSO.Check out your template. … use high quality Tag.
How can you tell if a primer has gel dimers?
The easiest way to check for primer-dimers is to compare your reactions to your negative control (water instead of DNA or RNA). Primer dimers will still form in the negative control. Some primer sets are more likely to form dimers than others.
What happens if primers are too long?
However, a primer should not be too long (> 30-mer primers) or too short. Short primers produce inaccurate, nonspecific DNA amplification product, and long primers result in a slower hybridizing rate. … One also needs to avoid primer-primer annealing which creates primer dimers and disrupts the amplification process.
How can primer dimer formation be avoided?
Prerequisites for avoiding primer-dimer formation during PCR include the design of optimal primer pairs, and the use of appropriate primer concentrations. … Reduce the primer concentration to the lowest amount at which product amplification can be achieved by conducting test runs with primer concentration gradients.How do you detect primer dimer in Qpcr?
The presence of primer dimers can be easily identified in the dissociation curve by the additional peak(s). The primer dimer peaks are usually seen at low melting temperatures. Use an amount of primer that produces no primer dimer and gives optimal amplification efficiency.
Can you run primers on a gel?
Agarose gel is not the best option for quality assurance of primers. Primers are too small for retardation by agarose, therefore denaturating acrylamide gel is the method of choise.
What is self dimer?
Self-dimers (also called homo-dimers) occur when some portion of an oligonucleotide is complementary to itself, resulting in an oligonucleotide molecule that can hybridize to another oligonucleotide molecule of the exact same sequence.
What is false priming?
Answers and Replies It likely refers to sequences of DNA that are very similar to the primer sequence and can bind the primer during the PCR reaction. These sites can cause false positives in PCR-based tests or cause extra bands to show up in a PCR reaction.What causes primer dimers in Qpcr?
As the name implies, primers dimerize mainly due to complementarity – either due to self complementarity of a single primer or complementarity due to primers designed for opposite strands. … Also, minimize high GC content at the 3′ end of the primer, this stabilizes the primer binding.
What does it mean if you have a band for your negative control?Whenever you see bands for your negative control, it generally means there was some form of contamination as Tomas has suggested. However, it does depend on the size of the bands. If the bands for the negative control show products much smaller than the samples or positive control, it could possibly be primer dimer.
Article first time published onWhat happens if you add too much primer to a PCR?
Too high primer concentrations increase the chance of mispriming, which results in nonspecific PCR products. Limiting primer concentrations result in extremely inefficient PCR reactions. The primer concentration can be calculated as described in Preparation of oligo solutions. DNA template concentration.
Are primers complementary to DNA?
Primers. – short pieces of single-stranded DNA that are complementary to the target sequence. The polymerase begins synthesizing new DNA from the end of the primer.
Does primer dimer affect qPCR?
the primer dimer effects the whole process of expression. As a matter of fact, existence of primer dimer decrease the synthesis of target gene and as a result you can not count on the expression pattern of your target gene. Most of qPCR have some level of primer dimer.
Where do primers anneal?
Annealing is usually done, at 5oC below the Tm of the primers, typically about 45-55oC. (As a simple rule of thumb, the Tm of the primers can be estimated by adding 2oC for each A or T and 4oC for each G or C.)
What causes nonspecific amplification PCR?
Extension time was too long: Excessive extension time can allow nonspecific amplification. Generally, use an extension time of 1 min/kb. 3. Annealing time was too long: Excessive annealing time may increase spurious priming.
How short can primers be?
A good length for PCR primers is generally around 18-30 bases. Specificity usually is dependent on length and annealing temperature. The shorter the primers are, the more efficiently they will bind or anneal to the target.
What is a PCR primer?
A primer is a short, single-stranded DNA sequence used in the polymerase chain reaction (PCR) technique. In the PCR method, a pair of primers is used to hybridize with the sample DNA and define the region of the DNA that will be amplified. Primers are also referred to as oligonucleotides.
What causes smearing in gel electrophoresis?
1. Improperly prepared gel: If the gel is not poured correctly, it will not polymerize or solidify evenly, thus causing the molecules to smear. … If the wells are filled too much, or if the sample is not properly diluted, the excess sample may smear across the gel.
What causes smearing in PCR?
DNA contamination, RNA in DNA sample, hight concentration of DNA in the PCR reaction can cause smearing. … DNA smearing usually caused in plants due to high concentration of template DNA.
How do you increase PCR yield?
- Check the primer design using computer software.
- Optimize the annealing temperature in a 1-2°C step.
- A primer concentration of 0.2 μM is satisfactory for most PCR reactions. …
- Increase cycling numbers up to 45 cycles.
- Do a manual hot-start.
- Use thin-wall 0.2 ml PCR tubes.
Do primer dimers affect sequencing?
Adapter dimers contain full-length adapter sequences that are able to bind and cluster on the flow cell and generate sequencing data. In contrast, primer dimers do not contain complete adapter sequences, and are not able to bind or cluster on the flow cell, so are not sequenced.
What is NTC in Qpcr?
A no template control (NTC) omits any DNA or RNA template from a reaction, and serves as a general control for extraneous nucleic acid contamination. … A no amplification control (NAC) omits the DNA polymerase from the PCR reaction.
What do dimers do?
For example, dimers form in the cell membrane, where tyrosine-kinase receptors reside, and in the cytosol that contains microtubules composed of tubulin. In the nucleus, hormone receptors, acting as transcription factors, form dimers to increase stability and improve binding to DNA.
What is false priming in PCR?
Nonspecific-or “false”-priming may occur during first strand synthesis due to reverse transcriptase reactions primed by hairpin structures of the target RNA (Fig. 4). In addition, antisense and sense transcripts can act to prime their comple- mentary strand during first strand synthesis (16).
How can PCR contamination be prevented?
Separating pre- and post-amplification areas is key to preventing contamination. Prepare your PCR master mix in a template-free room (see next bullet), using reagents that never come into contact with potential sources of contamination. Maintain a separate area for analyzing PCR amplicons.
How many primers can be used in multiplex PCR?
In multiplex PCR, two or more primer sets designed for amplification of different targets are included in the same PCR reaction. Using this technique, more than one target sequence in a clinical specimen can be amplified in a single tube.
How do you get rid of PCR contamination?
- 1) Introduction.
- 2) Laboratory construction.
- 3) Unidirectional Workflow.
- 4) Pipetting Technique.
- 5) Frequently Changing Gloves.
- 6) Aseptic Cleaning Technique.
- 7) Include controls in your protocol.
Why is it called real time PCR?
In order to robustly detect and quantify gene expression from small amounts of RNA, amplification of the gene transcript is necessary. … This measurement is made after each amplification cycle, and this is the reason why this method is called real time PCR (that is, immediate or simultaneous PCR).
How do you optimize primer concentration in PCR?
Design both primers to have melting temperatures within 3°C of each other to simplify your PCR optimization. End with a G or C. Capping the 3′ end of your primer sequence with a G or C will strengthen primer annealing at the site of extension. Remember to add spacers for restriction enzyme cloning/isothermal assembly.
Why might it be important that the two primers used in a PCR reaction have very similar melting temperatures?
Primer melting temperature (Tm) – All primers in the reaction must have similar melting temperature (Tm) so they anneal to and dissociate from complementary DNA sequences at approximately the same temperatures, allowing each amplification to precede at the selected temperature.
How do you avoid multiple bands in PCR?
- do the reaction with a negative control (no template). …
- Increase the annealing temperature. …
- Redesign the primers and make the 3′ longer. …
- Increase annealing time if the non-specific products are shorter than your target. …
- Use less DNA template.
- Try touch-down PCR.