How does the process of gel electrophoresis separate DNA fragments
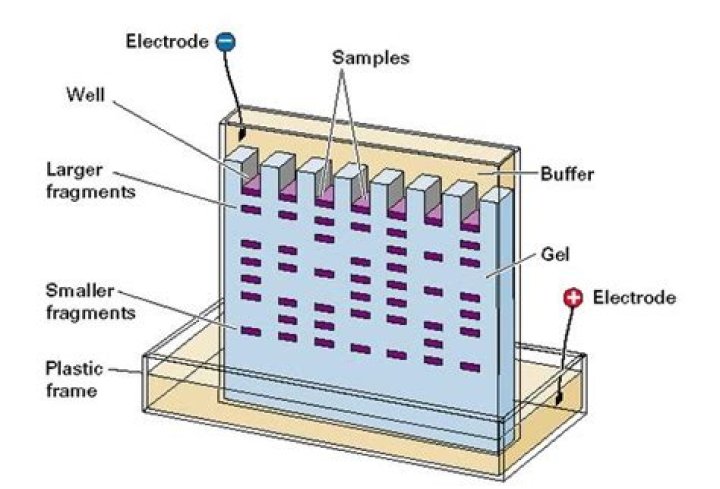
Gel electrophoresis is a technique used to separate DNA fragments according to their size. … DNA fragments are negatively charged, so they move towards the positive electrode. Because all DNA fragments have the same amount of charge per mass, small fragments move through the gel faster than large ones.
How does the process of gel electrophoresis separate DNA fragments quizlet?
How does the process of gel electrophoresis separate DNA fragments? It uses an electric current to separate different sized molecules of DNA in a porous sponge-like matrix. … Smaller fragments move faster, and therefore further, than larger fragments as they snake through the gel.
How does gel electrophoresis work quizlet?
How does gel electrophoresis work? Molecules are forced across a span of gel. Electrodes at either end of the gel provide the driving force. The charged particles migrate either to the cathode or to the anode.
What process will you use to separate the DNA fragments quizlet?
What process will you use to separate the DNA fragments? The restriction enzymes will now be separated by size-using a process known as gel electrophoresis. What does electrophoresis mean ?Why does the DNA separate in agarose gel electrophoresis?
Because DNA has a uniform mass/charge ratio, DNA molecules are separated by size within an agarose gel in a pattern such that the distance traveled is inversely proportional to the log of its molecular weight(3).
Why does gel electrophoresis work on DNA quizlet?
What does Gel Electrophoresis basically do to DNA? It separates the DNA into fragments, where electricity is run through the gel and the negatively charged DNA travels (the larger fragments traveling slower) towards the positive end of the gel.
What actually moves the DNA through the gel in gel electrophoresis?
Gel electrophoresis and DNA DNA is negatively charged, therefore, when an electric current is applied to the gel, DNA will migrate towards the positively charged electrode. Shorter strands of DNA move more quickly through the gel than longer strands resulting in the fragments being arranged in order of size.
What is the criterion for DNA fragments movement on agarose gel during gel electrophoresis?
What is the criterion for DNA fragments movement on agarose gel during gel electrophoresis? Explanation: During gel electrophoresis, DNA fragments move on an agarose gel according to size through the sieving effect. The smaller fragments move the farthest.How does gel electrophoresis work explain the process?
The process of gel electrophoresis works because negatively charged molecules move away from the negative pole of the electric current and smaller molecules will move faster than larger molecules. Thus, a size separation is achieved within the pool of molecules running through the gel.
How an agarose gel can separate DNA fragments of different lengths?The negatively charged DNA can be pulled toward the positive field of the gel. … Explain how an agarose gel can separate DNA fragments of different lengths. Smaller fragments move faster, and therefore further, than larger fragments as they snake through the gel.
Article first time published onWhat is the criterion for DNA fragments movement?
The larger the fragment size, the farther it moves.
What causes DNA fragments to migrate through the gel towards the anode?
Gel electrophoresis uses electricity to separate fragments of DNA based on their length. … The negative charge on the sugar-phosphate backbone of DNA polymers cause them to migrate towards the positive electrode when placed in an electrical field.
Which of the following technique is used for the separation of DNA fragments?
By using the agarose gel electrophoresis technique the DNA fragments can be separated.
Why is the DNA sample to be separated by gel electrophoresis always loaded at the cathode or negative end of the power source?
Why is the DNA sample to be separated by gel electrophoresis always loaded at the cathode or negative end of the power source? The gel acts as a molecular sieve: because nucleic acid molecules carry negative charges on their phosphate groups, they all travel toward the positive pole in an electric field.
How are DNA fragments separate on an agarose gel can be visualized?
In agarose gel electrophoresis, separated DNA fragments can be visualised with the help of ethidium bromide in UV radiation.
Which of the following is true regarding DNA fragments during gel electrophoresis?
“Which of the following is correct regarding the separation of DNA fragments during gel electrophoresis?” Smallest fragment will move to the farthest point towards cathode. Smallest fragment will move to the farthest point towards anode. … Largest fragment will move to the farthest point towards cathode.
What are fragments of DNA?
DNA fragmentation is the separation or breaking of DNA strands into pieces. It can be done intentionally by laboratory personnel or by cells, or can occur spontaneously. Spontaneous or accidental DNA fragmentation is fragmentation that gradually accumulates in a cell.
Which of the following is commonly used as a vector for introducing a DNA fragment?
Retrovirus is commonly used as vector for introducing a DNA fragment in human Iymphocyte.
Which vector can clone only a small fragment of DNA?
Plasmid is the vector which clones only a small fragment of DNA. Plasmids are autonomously replicating circular extra-chromosomal DNA. They are the standard cloning vectors and the ones most commonly used.
Which of the following technique is most commonly used to separate DNA?
The correct answer is (b) electrophoresis. Electrophoresis is the process of separating DNA molecules base on their size.
Which technique can be used to separate DNA fragments generated by restriction endonuclease?
The DNA fragments produced by restriction endonuclease digestion can be separated by gel electrophoresis, typically using agarose gels, and then visualized by staining with ethidium bromide.
Which technique is most suitable for the separation of large DNA fragments?
Agarose gel electrophoresis is the technique that is best suited for the separation of large DNA fragments.