How is Minisatellite DNA used
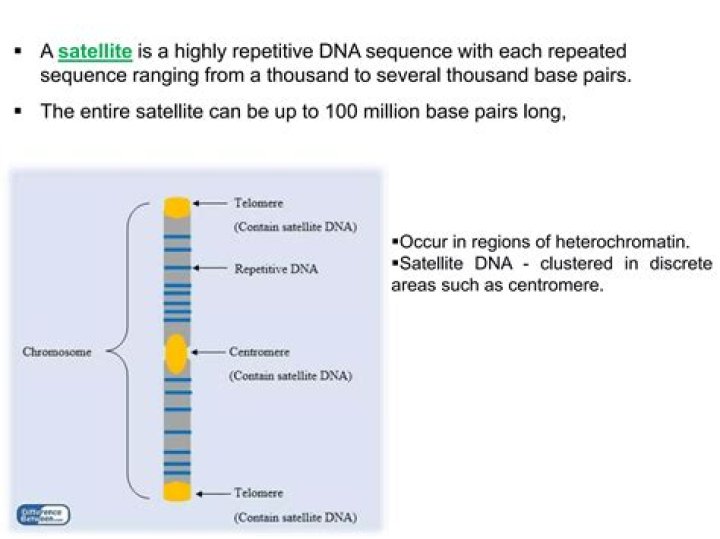
Minisatellites are the most highly variable sequence element in the human genome and the variable number of tandem repeats
Why is minisatellite DNA used for DNA fingerprinting?
Minisatellite alleles up to 5-10 kb long can be faithfully amplified. At least six minisatellite loci can be co-amplified from the same DNA sample and simultaneously detected to provide a reproducible and highly variable DNA fingerprint which can be obtained from nanogram quantities of human DNA.
What is the difference between minisatellite and microsatellite DNA?
Minisatellite is a section of highly repeated DNA that consists of a series of a repeating sequence composed of 10 to 100 base pairs. Microsatellite is a section of repetitive DNA that consists of short repeating sequences composed of 1 to 9 base pairs.
What are VNTRs useful for?
VNTRs are an important source of RFLP genetic markers used in linkage analysis (mapping) of genomes. They have become essential in forensic crime investigations. … Therefore, VNTRs are being used to study genetic diversity (DNA fingerprinting) and breeding patterns in animals.What is an example of mini satellite DNA?
Tandem repeats are repeated nucleotide sequences in which the copies lie adjacent to each other. It may be repetition(s) of one or more nucleotides. For example, CG CG CG CG CG is a tandem repeat wherein the sequence CG is repeated five times.
What is minisatellite 12 Mcq?
Hint:Minisatellites are tracts of DNA nucleotides consisting of around 10-60 base pairs long which are repeated several times and are found at numerous locations in the human genome.
What is a minisatellite DNA?
Minisatellite DNA, sometimes called variable number tandem repeats (VNTRs), is composed of blocks of longer repeats also dispersed throughout the genome. There is no known function for satellite DNA, nor is it known how the repeats are created.
Why VNTRs are considered as key factor in DNA profiling?
Variable number of tandem repeats are the short nucleotide sequences. These differ vary among all the individuals and make their DNA unique from other organism. VNTRs are the key factor in DNA profiling as they are not similar among the organisms. Thus, the correct answer is option (b).How VNTR is used in paternity testing?
Paternity testing using DNA polymorphism of variable numbers of tandem repeat (VNTR) regions with restriction fragment length polymorphism (RFLP) was implemented. … The comparison of 1,624 DNA fragments from 342 mother/child pairs showed only one difference above 1.25 mm which was interpreted as a mutation.
What are VNTRs mention the steps to detect VNTRs in identifying criminals in forensic investigations?Mention the steps to detect VNTR’s in identifying criminals in forensic investigations.” The DNA fragments that shows very high degree polymorphism. (ii) Digestion of DNA by restriction endonucleases. (iii) separation of DNA fragments by electrophoresis.
Article first time published onWhat range Minisatellite DNA units are repeated per repeat of minisatellite DNA in human genome?
The repeat unit of minisatellite DNA (also named VNTR previously) usually ranges from 10 to 100 bp and the block often extends between 100 bp and 20 kb.
Which satellite DNA is used in DNA fingerprinting?
Modern-day DNA profiling is also called STR analysis and relies on microsatellites rather than the minisatellites used in DNA fingerprinting. Microsatellites, or short tandem repeats (STRs), are the shorter relatives of minisatellites usually two to five base pairs long.
Does genome include RNA?
A genome is the complete set of DNA (or RNA in RNA viruses) of an organism. It is sufficient to build and maintain that organism. … The genome includes both coding regions (genes) and non-coding DNA, probably present in the nucleus, mitochondrion, chloroplast (for plants), and cytoplasm.
What is a minisatellite profile?
A minisatellite is a tract of repetitive DNA in which certain DNA motifs (ranging in length from 10 to 60 base pairs) are typically repeated 5–50 times. Minisatellites are notable for their high mutation rate and high diversity in the population, and they occur at more than 1000 locations in the human genome.
What are microsatellite markers?
Microsatellite markers are co-dominant, polymorphic DNA loci containing repeated nucleotide sequences, typically with 2 to 10 nucleotides per repeated unit.
Are Minisatellites coding?
Function. Minisatellites have been implicated as regulators of gene expression (e.g. at levels of transcription, alternative splicing, or imprint control). They are generally non-coding DNA but sometimes are part of possible genes.
Who discovered Minisatellite DNA?
1. The utilization of polymorphic DNA markers, minisatellites (variable number tandem repeats), and microsatellites [short tandem repeats (STRs)] for human identification in forensic genetics was originally proposed by Sir Alec Jeffreys, University of Leicester, United Kingdom.
Which technique is used with DNA fingerprinting Mcq?
Which of the following term is associated with DNA finger printing? Explanation: RFLP technique is preferred in forensic laboratories to identify the suspects.
What is VNTRs 12?
Hint: VNTRs are small DNA fragments which are 15-100 base pairs in length. They are repeating DNA strands which are found within and between the genes. These are found on the non-coding part of the genome and are used in VNTR profiling.
How is VNTR different from probe?
VNTR stands for Variable Number of Tandem Repeats. Probe is labelled or radioactive single stranded polynucleotide that hybridises with DNA fragments. Whereas, VNTR is a class of sateltite that shows high degree of polymorphism.
How do VNTRs differ?
While the repeated sequences themselves are usually the same from person to person, the number of times they are repeated tends to vary. … VNTRs are similar to Short Tandem Repeats (For more on STRs, see page 3), the difference being that in a VNTR, the repeated sequence is longer — about 10-100 base pairs long.
Is VNTR a Minisatellite?
VNTRs are a type of minisatellite in which the size of the repeat sequence is generally ten to one hundred base pairs. Minisatellites are a type of DNA tandem repeat sequence, meaning that the sequences repeat one after another without other sequences or nucleotides in between them.
Why must you use restriction enzymes in a DNA fingerprinting experiment?
– restriction enzymes are so significant in the process of DNA Fingerprinting because, in order to be able to sequence DNA, a method of cutting the DNA molecule into smaller fragments at precise locations is necessary.
What are tandemly arranged repeats?
= A tandem repeat is a sequence of two or more DNA base pairs that is repeated in such a way that the repeats lie adjacent to each other on the chromosome. Tandem repeats are generally associated with non-coding DNA. In some instances, the number of times the DNA sequence is repeated is variable.
How many microsatellites are in the human genome?
Each microsatellite consist of a short motif (1–6 base pairs) repeated in tandem to form an array [2]; over 600,000 unique microsatellites exist in the human genome [3, 4].
What are some applications uses of DNA profiling?
- identify the probable origin of a body fluid sample associated with a crime or crime scene.
- reveal family relationships.
- identify disaster victims, for example, ESR scientists travelled to Thailand to help identify victims of the 2004 Boxing Day tsunami.
How is DNA fingerprinting used?
DNA fingerprinting is a laboratory technique used to establish a link between biological evidence and a suspect in a criminal investigation. A DNA sample taken from a crime scene is compared with a DNA sample from a suspect. If the two DNA profiles are a match, then the evidence came from that suspect.
What are five other uses of DNA fingerprinting?
- establish paternity and parentage.
- identify victims of war and large scale disasters.
- study biodiversity of species.
- track genetically modified crops.
- settle immigration disputes.
What does a genome do?
A genome is the complete set of genetic information in an organism. It provides all of the information the organism requires to function. In living organisms, the genome is stored in long molecules of DNA called chromosomes.
What is the difference between a gene and a genome?
A gene consists of enough DNA to code for one protein, and a genome is simply the sum total of an organism’s DNA. DNA is long and skinny, capable of contorting like a circus performer when it winds into chromosomes.
What is genome sequencing Covid?
Genome sequencing for COVID-19 is about developing a complete picture of a virus’s RNA. It involves obtaining positive COVID-19 samples and generating a complete RNA sequence of that virus from that sample.



